_PATHWAY_93.png)
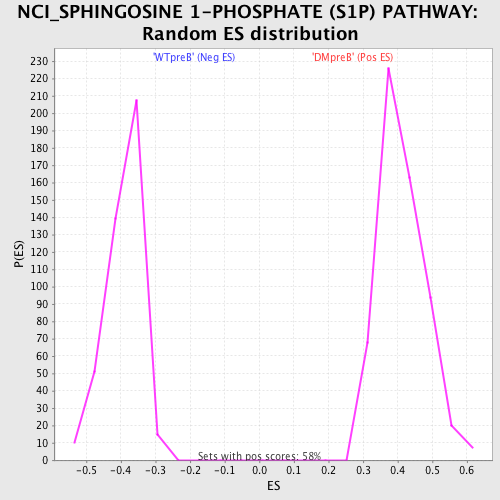
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | NCI_SPHINGOSINE 1-PHOSPHATE (S1P) PATHWAY |
| Enrichment Score (ES) | -0.6666101 |
| Normalized Enrichment Score (NES) | -1.7094414 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.010424511 |
| FWER p-Value | 0.089 |
_PATHWAY_93.png)
| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | BAX | 17832 | 644 | 4.463 | -0.0074 | No | ||
| 2 | CYCS | 8821 | 744 | 4.120 | 0.0125 | No | ||
| 3 | CTSD | 22 | 994 | 3.332 | 0.0194 | No | ||
| 4 | MAPK8 | 6459 | 1644 | 2.061 | -0.0030 | No | ||
| 5 | BID | 17017 435 | 1954 | 1.590 | -0.0100 | No | ||
| 6 | BAG4 | 7431 | 2121 | 1.411 | -0.0103 | No | ||
| 7 | BCAR1 | 18741 | 2225 | 1.277 | -0.0080 | No | ||
| 8 | PRKCZ | 5260 | 2281 | 1.203 | -0.0036 | No | ||
| 9 | PAWR | 19885 658 | 2446 | 1.028 | -0.0062 | No | ||
| 10 | HRAS | 4868 | 2610 | 0.906 | -0.0094 | No | ||
| 11 | SLC9A3R2 | 23089 1544 1514 | 3024 | 0.543 | -0.0284 | No | ||
| 12 | ITGB3 | 20631 | 3043 | 0.531 | -0.0262 | No | ||
| 13 | VEGFA | 22969 | 3350 | 0.369 | -0.0404 | No | ||
| 14 | FADD | 17536 8950 4711 | 3666 | 0.282 | -0.0557 | No | ||
| 15 | GNAI1 | 9024 | 3780 | 0.255 | -0.0603 | No | ||
| 16 | SHC1 | 9813 9812 5430 | 3936 | 0.225 | -0.0673 | No | ||
| 17 | GNAQ | 4786 23909 3685 | 4136 | 0.194 | -0.0769 | No | ||
| 18 | AKT1 | 8568 | 4218 | 0.183 | -0.0801 | No | ||
| 19 | GNAZ | 71 | 4228 | 0.182 | -0.0795 | No | ||
| 20 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 4380 | 0.162 | -0.0867 | No | ||
| 21 | FYN | 3375 3395 20052 | 5072 | 0.109 | -0.1233 | No | ||
| 22 | SNX15 | 23993 3761 | 5222 | 0.100 | -0.1308 | No | ||
| 23 | SPHK1 | 5484 1266 1402 | 5473 | 0.088 | -0.1438 | No | ||
| 24 | SRC | 5507 | 5727 | 0.076 | -0.1570 | No | ||
| 25 | IGF1 | 3352 9156 3409 | 6411 | 0.055 | -0.1936 | No | ||
| 26 | GNAO1 | 4785 3829 | 6926 | 0.044 | -0.2211 | No | ||
| 27 | PTGS2 | 5317 9655 | 7102 | 0.040 | -0.2303 | No | ||
| 28 | STAT5B | 20222 | 7325 | 0.036 | -0.2421 | No | ||
| 29 | PAK1 | 9527 | 7590 | 0.031 | -0.2562 | No | ||
| 30 | SHB | 10493 | 7714 | 0.029 | -0.2626 | No | ||
| 31 | HIF1A | 4850 | 7746 | 0.029 | -0.2641 | No | ||
| 32 | NCK1 | 9447 5152 | 7819 | 0.028 | -0.2679 | No | ||
| 33 | KDR | 16509 | 8027 | 0.024 | -0.2789 | No | ||
| 34 | GRB10 | 4799 | 8036 | 0.024 | -0.2792 | No | ||
| 35 | PDGFB | 22199 2311 | 8207 | 0.022 | -0.2882 | No | ||
| 36 | GNA14 | 23907 | 8214 | 0.022 | -0.2884 | No | ||
| 37 | RHOA | 8624 4409 4410 | 8261 | 0.021 | -0.2908 | No | ||
| 38 | PTPN2 | 23414 | 8306 | 0.020 | -0.2930 | No | ||
| 39 | GNA12 | 9023 | 8597 | 0.016 | -0.3086 | No | ||
| 40 | ACP1 | 4317 | 9230 | 0.009 | -0.3427 | No | ||
| 41 | PTPN11 | 5326 16391 9660 | 9360 | 0.007 | -0.3497 | No | ||
| 42 | FLT1 | 3483 16287 | 9989 | -0.001 | -0.3836 | No | ||
| 43 | IRS1 | 4925 | 10290 | -0.005 | -0.3998 | No | ||
| 44 | CDC42 | 4503 8722 4504 2465 | 10407 | -0.006 | -0.4060 | No | ||
| 45 | LYN | 16281 | 10463 | -0.007 | -0.4090 | No | ||
| 46 | CRADD | 19640 | 10936 | -0.013 | -0.4344 | No | ||
| 47 | YES1 | 5930 | 10960 | -0.014 | -0.4356 | No | ||
| 48 | LRP1 | 9284 | 11848 | -0.027 | -0.4833 | No | ||
| 49 | PLCB2 | 5262 | 11963 | -0.029 | -0.4893 | No | ||
| 50 | DOK1 | 17104 1018 1177 | 12045 | -0.031 | -0.4935 | No | ||
| 51 | EGF | 15169 | 12130 | -0.033 | -0.4978 | No | ||
| 52 | PIK3R1 | 3170 | 12502 | -0.041 | -0.5177 | No | ||
| 53 | PLCG1 | 14753 | 12675 | -0.046 | -0.5267 | No | ||
| 54 | SOS1 | 5476 | 13046 | -0.056 | -0.5463 | No | ||
| 55 | GNAI3 | 15198 | 13219 | -0.062 | -0.5552 | No | ||
| 56 | SPHK2 | 1217 1806 12155 | 13302 | -0.065 | -0.5593 | No | ||
| 57 | PDGFRB | 23539 | 13453 | -0.070 | -0.5670 | No | ||
| 58 | PIK3CA | 9562 | 13540 | -0.075 | -0.5711 | No | ||
| 59 | ASAH1 | 3835 3853 8630 | 13543 | -0.075 | -0.5708 | No | ||
| 60 | TNF | 23004 | 13668 | -0.082 | -0.5770 | No | ||
| 61 | ABL1 | 2693 4301 2794 | 14050 | -0.106 | -0.5969 | No | ||
| 62 | GRB7 | 20673 | 14124 | -0.112 | -0.6002 | No | ||
| 63 | PDGFA | 5237 | 14395 | -0.140 | -0.6139 | No | ||
| 64 | CBL | 19154 | 14602 | -0.166 | -0.6241 | No | ||
| 65 | FOS | 21202 | 14800 | -0.201 | -0.6335 | No | ||
| 66 | SMPD3 | 18757 | 14820 | -0.203 | -0.6333 | No | ||
| 67 | MAP2K2 | 19933 | 14908 | -0.221 | -0.6366 | No | ||
| 68 | GNA15 | 19671 | 14942 | -0.226 | -0.6370 | No | ||
| 69 | VAV2 | 5848 2670 | 14978 | -0.236 | -0.6375 | No | ||
| 70 | JAK2 | 23893 9197 3706 | 15111 | -0.267 | -0.6430 | No | ||
| 71 | STAT5A | 20664 | 15113 | -0.268 | -0.6414 | No | ||
| 72 | AIFM1 | 2558 24161 | 15116 | -0.268 | -0.6398 | No | ||
| 73 | MAP2K1 | 19082 | 15318 | -0.325 | -0.6487 | No | ||
| 74 | MYC | 22465 9435 | 15334 | -0.330 | -0.6475 | No | ||
| 75 | LCK | 15746 | 15440 | -0.368 | -0.6509 | No | ||
| 76 | TNFRSF1A | 1181 10206 | 15447 | -0.370 | -0.6490 | No | ||
| 77 | CDH5 | 18512 | 15470 | -0.378 | -0.6478 | No | ||
| 78 | FGR | 4723 | 15758 | -0.526 | -0.6601 | No | ||
| 79 | MAP4K4 | 11241 | 15849 | -0.588 | -0.6614 | No | ||
| 80 | ITGAV | 4931 | 15941 | -0.673 | -0.6622 | Yes | ||
| 81 | HCK | 14787 | 15979 | -0.695 | -0.6599 | Yes | ||
| 82 | RIPK1 | 5381 | 16037 | -0.757 | -0.6584 | Yes | ||
| 83 | PTPN1 | 5325 | 16168 | -0.897 | -0.6599 | Yes | ||
| 84 | TRAF2 | 14657 | 16252 | -0.981 | -0.6584 | Yes | ||
| 85 | PTPRJ | 9664 | 16387 | -1.089 | -0.6589 | Yes | ||
| 86 | NCK2 | 9448 | 16530 | -1.250 | -0.6589 | Yes | ||
| 87 | RAC1 | 16302 | 16651 | -1.434 | -0.6566 | Yes | ||
| 88 | BIRC3 | 19247 | 16725 | -1.544 | -0.6511 | Yes | ||
| 89 | AKT3 | 13739 982 | 16769 | -1.612 | -0.6436 | Yes | ||
| 90 | RASA1 | 10174 | 16815 | -1.689 | -0.6356 | Yes | ||
| 91 | MAPK3 | 6458 11170 | 16931 | -1.809 | -0.6308 | Yes | ||
| 92 | SMPD1 | 18145 2658 | 16969 | -1.861 | -0.6213 | Yes | ||
| 93 | CASP8 | 8694 | 17013 | -1.927 | -0.6119 | Yes | ||
| 94 | MAP2K4 | 20405 | 17051 | -1.981 | -0.6017 | Yes | ||
| 95 | RAF1 | 17035 | 17131 | -2.125 | -0.5929 | Yes | ||
| 96 | SGPL1 | 19757 | 17239 | -2.305 | -0.5846 | Yes | ||
| 97 | PRKCD | 21897 | 17290 | -2.395 | -0.5726 | Yes | ||
| 98 | MAPK14 | 23313 | 17421 | -2.682 | -0.5632 | Yes | ||
| 99 | BAD | 24000 | 17524 | -2.914 | -0.5508 | Yes | ||
| 100 | SGPP1 | 21039 | 17527 | -2.920 | -0.5330 | Yes | ||
| 101 | GRB2 | 20149 | 17590 | -3.055 | -0.5177 | Yes | ||
| 102 | PRKRA | 357 | 17594 | -3.069 | -0.4990 | Yes | ||
| 103 | JUN | 15832 | 17687 | -3.335 | -0.4835 | Yes | ||
| 104 | GNA13 | 20617 | 17715 | -3.402 | -0.4641 | Yes | ||
| 105 | SLC9A3R1 | 11250 | 17768 | -3.541 | -0.4452 | Yes | ||
| 106 | NFKB1 | 15160 | 17789 | -3.591 | -0.4243 | Yes | ||
| 107 | GNA11 | 3300 3345 19670 | 17835 | -3.697 | -0.4041 | Yes | ||
| 108 | NFKBIA | 21065 | 17838 | -3.699 | -0.3815 | Yes | ||
| 109 | RELA | 23783 | 17959 | -4.092 | -0.3629 | Yes | ||
| 110 | PTEN | 5305 | 18200 | -4.996 | -0.3452 | Yes | ||
| 111 | GAB1 | 18828 | 18283 | -5.430 | -0.3164 | Yes | ||
| 112 | STAT1 | 3936 5524 | 18294 | -5.505 | -0.2832 | Yes | ||
| 113 | CSK | 8805 | 18359 | -5.871 | -0.2506 | Yes | ||
| 114 | BLK | 3205 21791 | 18378 | -5.983 | -0.2149 | Yes | ||
| 115 | MAP3K1 | 21348 | 18411 | -6.282 | -0.1781 | Yes | ||
| 116 | STAT3 | 5525 9906 | 18455 | -6.760 | -0.1390 | Yes | ||
| 117 | DNM2 | 4635 3103 | 18471 | -6.991 | -0.0969 | Yes | ||
| 118 | CXCR4 | 13844 | 18551 | -7.968 | -0.0524 | Yes | ||
| 119 | MAPK1 | 1642 11167 | 18592 | -9.104 | 0.0013 | Yes |
_PATHWAY_94.png)